the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Heterogeneity of the urban soil microbiome and associations with physicochemical soil characteristics
Taline Bosiers
Nona De Groof
Esther Diels
Jente Dubois
Fine Gilis
Agustina Santullo Latorre
Karen Wuyts
Urban soil plays a fundamental role in supporting a range of ecosystem services important for both human health and urban resilience. While many ecosystem services rely on soil microbial communities, studies on urban soil microbes have remained scarce. A major complexity in understanding urban soil bacteria is the heterogeneity of soil throughout the city. In this study, we investigated the urban soil bacterial communities of street tree areas in relation to soil characteristics. We sampled soils under London plane trees at 20 street sites throughout the city of Antwerp (Belgium) and looked into the horizontal and vertical spatial variation in soil bacterial communities. Using 16S rRNA gene amplicon sequencing, we found great bacterial diversity (over 1800 bacterial genera) and heterogeneity in the urban soil of a single city. Our findings suggest that, first and foremost, urban soil community variation is determined by the sampling site, indicating that samples grouped together in space are more similar. Additionally, but to a smaller extent, the bacterial communities are affected by soil characteristics as is the case for non-urban soils. Significant relationships were observed between composition and soil pH, moisture, density, depth, air pollution, and land use class (all p values < 0.01). We found that soils, especially those with relatively low moisture content in the city under study, showed a decline in soil bacterial biodiversity with decreasing moisture content (p value = 0.047), indicating that soil moisture content may be an important aspect of sustainable urban soil management. Furthermore, soils under trees with a smaller circumference varied greatly in bacterial community composition relative to soils under trees with a bigger circumference, which were significantly more alike in bacterial composition. This indicates that, to better predict urban resilience, factors such as soil and vegetation development should be taken into account. In conclusion, we recommend including microbial soil diversity in urban sustainable soil management and focusing future urban soil research on understanding what the desired microbial functions and compositions for urban soils are.
- Article
(2372 KB) - Full-text XML
-
Supplement
(585 KB) - BibTeX
- EndNote
Urban soil plays a fundamental role in supporting a range of ecosystem services important for both human health and urban resilience. These include countering the urban heat island effect, mitigating floods, capturing and degrading pollutants, providing essential support for infrastructure, facilitating urban agriculture, and offering access to green spaces for mental and physical health (O'riordan et al., 2021). Additionally, at the global level, soil contributes to processes such as nutrient cycling and carbon storage (Wang and Goll, 2021). Many ecosystem services depend on soil microbial communities, which play a crucial role in nutrient cycling, organic matter decomposition, plant health, and overall soil ecosystem functioning (Nugent and Allison, 2022). Moreover, there is a growing body of evidence suggesting that the interaction with natural microbial biodiversity can have positive effects on human health, primarily by influencing the microbiome and immune system (Haahtela, 2019; Mills et al., 2017). The importance of healthy soils, defined as soils in good chemical, physical, and biological condition, is corroborated by the European Union Soil Strategy for 2030 (European Commission, 2021) to help us address the biggest global challenges.
The soil microbiome of natural environments and agricultural lands has been extensively studied, while studies on urban soil microbes have remained scarce (Nugent and Allison, 2022). Nevertheless, many soil characteristics that change with urbanization can be expected to impact soil microbial communities (Nugent and Allison, 2022). The process of urbanization involves construction, sealing, and other human activities, which significantly alters the physical, chemical, and biological characteristics of soils (Lehmann and Stahr, 2007). Urban development often leads to soil compaction due to heavy construction machinery and increased traffic. Compaction reduces pore spaces in the soil, limiting water infiltration and aeration, which affects soil microorganisms and plant growth. Furthermore, soil moisture content profoundly influences microbial activity and diversity (Brockett et al., 2012). Moist soils provide a favourable environment for microbial growth and metabolic processes. Conversely, waterlogged or excessively dry conditions can limit microbial activity, affecting soil functions like organic matter decomposition. Construction activities also include moving soil and leaving behind construction debris, making urban soils very heterogeneous (Lehmann and Stahr, 2007). Moreover, urban areas are associated with increased pollution from vehicular emissions, industrial activities, and improper waste disposal, leading to soil contamination with heavy metals, organic pollutants, and other harmful substances (Yu et al., 2019; Bezberdaya et al., 2022; Yuan et al., 2021). Pollutants associated with traffic on motorways may also greatly affect soil chemistry and its microbial communities (Yan et al., 2016; Lovett et al., 2000). Soil pH has previously been argued to be the main explanatory variable for the composition of bacterial communities of urban and non-urban soils (Yan et al., 2016; Fierer and Jackson, 2006). Changes in pH can alter microbial community structure, affecting nutrient cycling and soil health (Malik et al., 2018). Hence, the impact of urbanization on microbial soil communities is expected to be multifaceted.
Urbanization-induced soil changes have indeed been observed to alter microbial diversity and composition. It has been shown that the amount and diversity of fungi, including fungi that are beneficial to plants, decrease drastically with urbanization (Abrego et al., 2020; Bainard et al., 2011). A study covering 56 cities around the world found that the functions of urban soil microbes tend to lean towards more greenhouse gas emissions, faster nutrient cycling, and more human and plant pathogenicity compared to nearby natural soils (Delgado-Baquerizo et al., 2021). However, soils exclusively from urban green spaces were included in this study.
One of the greatest challenges of research on the urban soil microbiome is the heterogeneous nature of urban soils (Lehmann and Stahr, 2007). We therefore set up an exploratory soil microbiome study in a single city, examining established drivers of microbiomes of street tree soils together with the microbiome variation across various spatial distance levels (2 m to several kilometres), reflecting soil heterogeneity. For soil characteristics, we focused on soil pH; soil moisture content; soil density; and indicators of pollution, such as soil remanent magnetization (for metals) and atmospheric coarse particulate matter (PM10) and nitrogen oxide (NOx) concentrations (indicating vehicular emissions). To ensure consistency of the vegetation effect on soils, we selected sampling sites associated with a single tree species, the London plane (Platanus × acerifolia), which is abundantly found in the city. Tree circumference was measured as a proxy for tree age. We sampled soils in unsealed soil volumes next to paved streets. With this exploratory study, we aimed to better understand how soil heterogeneity caused by human activity and classic soil characteristics relate in driving the urban soil microbiome structure.
2.1 Study area and design
This study was conducted in the city of Antwerp, the second-largest city of Belgium (51°13′16.5′′ N, 4°23′58.4′′ E), which houses 545 000 inhabitants (2024) on 205 km2. Antwerp is located in the sandy Campine region: the upper mineral soils are dominated by sand or sandy loams, although clay soils are sporadic due to the city's location on the river banks of the river Scheldt (City of Antwerp, 2023). More background information on the city can be found in Muyshondt et al. (2022).
Throughout the city, we chose 20 sites where solitary London plane street trees (Platanus × acerifolia) were present in paved streets and where the soil in which the tree was rooted was unsealed and thus accessible for sampling. These sites represent the largest spatial distance level in this study and were between 100 m and 7 km apart. At each site, soil samples were collected under two different trees (Fig. 1). Trees of the same site were between 3 and 20 m apart, representing the second-largest spatial distance level in this study. At each tree, two orientations (north and south) were sampled, which were between 1 and 3 m apart, representing the smallest spatial distance level in this study. Since these samples were taken close to the same tree, we can assume that they have a similar history of soil manipulation. Finally, at each orientation under each tree at each site, the upper mineral soil was sampled at two soil depths: from 0 to 5 cm and from 10 to 15 cm. In total, 160 soils were sampled between 13 February and 31 March 2023.
2.2 Sample collection and measurements
Samples for bacterial analysis were taken with a steel soil auger (Eijkelkamp Soil and Water, diameter 2.5 cm) and spatula. The soil borer samples were transferred into small plastic bags and stored in a freezer (−18 °C) immediately upon arrival at the lab (within 10 h). After each sampling, the tools were first cleaned of soil residues, then disinfected with a 70 % ethanol-impregnated paper towel. Samples for physical, chemical, and magnetic analyses were taken with Kopecky rings (height 5 cm, diameter 5 cm). Soil from the Kopecky rings was transferred into a plastic jar for transport. These soil samples were weighed and dried upon arrival in the lab as described below. Tree height (m) was measured using the Vertex IV altimeter and Transponder T3 (Haglöf, Sweden). Trunk circumference (cm) was determined at breast height with a tape measure as an indication of tree age (Van Mensel et al., 2023).
2.3 Physical, chemical, and magnetic analyses
After determination of the soil wet weight with an electronic balance (Kern and Sohn; accuracy 0.2 g), the samples taken with the Kopecky rings (Eijkelkamp) were dried for 1 week in a drying oven (Memmert) at 50 °C, and the soil dry weight was determined. The gravimetric soil moisture content (%) was determined as the difference between the wet and dry weight divided by the dry weight of the Kopecky samples. The density (kg m−3) was determined from the dimensions of the Kopecky ring (volume 9.8 × 10−5 m3) and the dry weight. The pH was measured by mixing 10 g of dry soil in 25 mL of 1 M KCl (Wuyts et al., 2013).
The soil saturation isothermal remanent magnetization (SIRM) was measured as a proxy for exposure to traffic-derived pollution (Lu et al., 2011). For this magnetic analysis, the dried samples were sieved with a stainless steel sieve with a mesh size of 2 mm. A sub-sample of about 3 g was weighted and tightly wrapped in plastic film to prevent movement of the soil particles and placed in a 6.7 cm3 plastic sample container. The SIRM of this container was determined from the remanent magnetization (A m−1) measured with a JR-6 dual-speed spinner magnetometer (AGICO; sensitivity 2.4 × 10−6 A m−1; accuracy 3 %) after magnetization with a pulse magnetizer (Molspin) at 0.8 Tesla. Blanks were taken from the plastic sample container along with the same amount of plastic film used for wrapping the soil samples. For normalization of the SIRM (A m2 kg−1), the average blank was subtracted from the measured value, then divided by the dry weight of the sample, and finally multiplied by the volume of the sample container (6.7 cm3).
2.4 Molecular analysis
Soil samples for bacterial analysis were thawed and sieved with a stainless steel sieve with a mesh size of 2 mm right before DNA extractions were performed using the DNeasy PowerSoil Pro Kit (QIAGEN, Germany) according to the manufacturer's instructions. A PCR was then set up according to Wuyts et al. (2020). Blanks for DNA extraction and PCR were included. Amplified samples were purified with the Agencourt AMPure XP (Beckman Coulter, USA) according to the manufacturer's instructions. Finally, all samples were pooled at equimolar concentrations into a library. For this, the concentration of the individual samples was determined using the Qubit 3.0 fluorometer (ThermoFisher Scientific, USA). The library was sequenced at the Centre for Medical Genetics (Edegem, Belgium) using Illumina MiSeq. In total, 158 bacterial community samples were sequenced with an additional nine blanks, representing both DNA extraction kit blanks and PCR-negative controls.
2.5 Land use and air quality
Modelled atmospheric concentrations (µg m−3) of particulate matter with an aerodynamic diameter smaller than 10 µm (PM10) and nitrogen oxides (NOx) were used as indicators of air pollution. These were extracted for the different sites from air quality maps generated with the ATMO-Street model 2021 (http://ftp.irceline.be/atmostreet/, last access: 27 January 2026). Although previous research has shown that air pollution spatial variation can be large in urban settings and not fully accounted for in modelled values (Hofman et al., 2016), these model values provide the best available indication of air quality at the sites.
The predominant land use class within a 200 m radius around the sites was extracted using Urban Atlas 2018 (European Union's Copernicus Land Monitoring Service Information, 2020). All spatial analyses were performed via QGIS version 3.22.16 (http://qgis.org, last access: 27 January 2026). To reduce the number of underrepresented land use classes, the classes “industrial, commercial, public, military, and private units”, “fast transit roads and associated land”, and “other roads and associated land” were merged into one category, namely “roads and associated land”. The classes “discontinuous dense urban fabric” and “discontinuous medium-density urban fabric” were merged into the category “discontinuous urban fabric”.
2.6 Bioinformatics and statistical analyses
The raw sequencing data were processed using the DADA2 package (Callahan et al., 2016) in the R environment (R Core Development Team, 2006). In short, sequence reads with at least one ambiguous base, reads containing the lowest possible quality score of 2, and reads with more than two “expected errors” were discarded. Next, sample inference (DADA2's core algorithm) was performed, followed by read merging. A sequence table containing all amplicon sequence variants (ASVs) was constructed, chimeras were removed, and taxonomy was assigned using a custom Genome Taxonomy Database (GTDB; release 207) 16S rRNA sequence database, complemented with mitochondrial and chloroplast sequences from SILVA v138. Non-bacterial reads were removed before analyses. Inspection of the blanks indicated that the ASV Moraxella_A was a consistent contaminant, and after establishing its very low abundance in the samples, we removed it from our dataset. Furthermore, all six extraction kit blanks and three PCR blanks had very low to no read numbers (fewer than 32), indicating very low to no contamination, and were removed from the dataset before analyses. Finally, samples with fewer than 1000 reads were discarded.
Statistical analyses were done in the R environment (R Core Development Team, 2006). Graphs were generated using the ggplot2 package (Wickham, 2016) and ggbiplot (Vu et al., 2011). Whenever the p value was under 0.05, the result was considered significant. The natural logarithm of the SIRM value was used in further analyses because of the right-skewed distribution. Means are reported as mean ± standard deviation. For soil pH, medians were reported. For correlations between soil characteristics, the Pearson correlation was calculated. For correlations with factors that were determined per site (e.g. atmospheric PM10), the mean values per site were used. Four mixed-effect models were assessed using the lmerTest package (Kuznetsova et al., 2017) to test if variation in continuous soil properties (soil pH, soil moisture content, soil ln(SIRM), and soil density) can be explained by depth (two levels), orientation (two levels), and land use (three levels). Site was added to the model as a random effect. Additionally, a principal component analysis was done to visualize the correlations of all variables and cluster the sites in a two-dimensional space based on the soil variables (pH, moisture, and ln(SIRM)) and the environmental variables (i.e. atmospheric PM10 concentration, atmospheric NOx concentration, tree height, and stem circumference).
For bacterial community analyses, the tidytacos package (Wittouck et al., 2025) in R was used. Bray–Curtis dissimilarities of bacterial communities were calculated to perform permutational multivariate analysis of variance (PERMANOVA) using the adonis2 function (Oksanen et al., 2022). We tested two PERMANOVA models with the complete community dataset: once using soil characteristics to explain community variation and once using factors related to spatial distance level to explain community variation. For the latter, the largest spatial distance level, site, was put first in the model. The next spatial distance level was tree ID, which was nested in site, and the following factors, depth and orientation, were nested within tree ID. In addition to PERMANOVA, we used PERMDISP2 of the vegan package (Oksanen et al., 2022) to test for multivariate homogeneity of groups using the distance from each data point to its group centroid and then testing whether those distances differed among the groups defined by the variables. To further study the interaction effects in the PERMANOVA model with soil characteristics, we made four subsets of our dataset. At each depth, we merged the bacterial samples of each tree and determined median pH and ln(SIRM) values per tree. To study the interaction effect with tree circumference, we then ran PERMANOVA models on four subsets to assess how the impact of pH and SIRM changes between low (< 100 cm) and high (> 100 cm) trunk circumference at the two different soil depths.
We also tried to detect ecological succession and community convergence of the bacterial communities over time as described by Shade et al. (2013) and Smets et al. (2022). We furthermore determined alpha diversity of the bacterial communities. Therefore, the dataset was first rarefied to 1140 reads, and the observed ASV richness and ASV inverse Simpson index were then calculated. Correlations between these two diversity metrics and the soil variables were determined using generalized linear models which included the total read counts per sample. For pH we also tested a quadratic regression model, as it was expected to show an optimum.
3.1 Soil environmental properties
Several soil, air, and tree properties were recorded for each sample in this study and are summarized in Table 1. Correlation analyses (Table 2, Fig. 2) showed a negative correlation between soil moisture content and soil pH, while moisture content was positively correlated with soil ln(SIRM) value. In addition, there was a significant positive correlation between atmospheric PM10 concentrations and atmospheric NOx concentrations.

Figure 2PCA of the different soil and environmental properties per soil sample recorded in this study. The colour of the dots indicates (a) the soil depth of where the sample was taken, (b) the orientation of where the sample was taken compared to the focal tree, and (c) the major land use class of the site where the sample was taken.
Table 1The mean, standard deviation, and range of the soil, atmospheric, and tree properties recorded in this study.
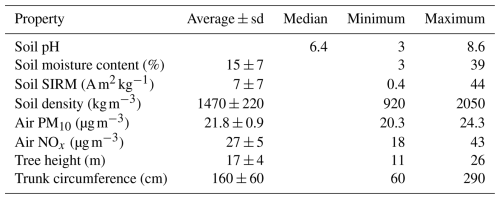
Table 2Correlations between environmental variables, including soil properties and air quality.
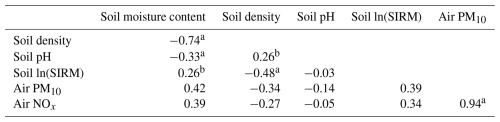
a p value < 0.001, b p value < 0.01.
A mixed-effect model showed that variation in soil moisture content was significantly explained by depth (p < 0.001), orientation (p = 0.024), and land use (p = 0.025). The average moisture content at a depth of 0–5 cm (17 % ± 8 %) was higher than at a depth of 10–15 cm (12 % ± 4 %) and higher at the northern orientation (16 % ± 7 %) than at the southern orientation (14 % ± 7 %). Furthermore, the average moisture content was higher in the “discontinuous urban fabric” land use class (18 % ± 7 %) compared to the “continuous urban fabric” (sealing > 80 %) land use class (13 % ± 5 %).
Another mixed-effect model showed that there was a significant difference in soil pH at the different orientations (p = 0.033) and land uses (p = 0.048). The median pH at the southern orientation was 6.54, while it was 6.24 at the northern orientation. The median pH for land use classes “roads and associated land”, “discontinuous urban fabric”, and “continuous urban fabric” (sealing > 80 %) was 5.72, 5.95, and 6.75, respectively. The variation in soil SIRM and soil density were not significantly explained by depth, orientation, or land use.
The principal component analysis (PCA) does not show particularly obvious clustering of the sampling sites (Fig. 2), but most of the variation between soil samples can be found in atmospheric concentrations together with soil SIRM on the one hand (principal component 1, PC1) and in soil pH together with soil density in a negative relationship with soil moisture on the other hand (principal component 2, PC2). PC1 and PC2 explain 59 % of the total variation in soil, atmospheric, and tree characteristics.
3.2 Relationships between soil characteristics and bacterial communities
After quality filtering of the DNA sequencing data as described in the methods, the resulting dataset contained approximately 1.3 million reads of 122 soil samples with 11 093 amplicon sequence variants (ASVs), representing 1835 genera. The overall most abundant ASVs belonged to the genera Renibacterium (2.8 %), Bradyrhizobium (1.5 %), Nitrosocosmicus (1.1 %), and Hyphomicrobium (1.1 %). The most abundant phyla in our dataset were Proteobacteria (24 %), Acidobacteriota (20 %), Actinobacteriota (19 %), Firmicutes (7 %), and Verrucomicrobiota (6 %). The raw sequencing data are publicly available on the European Nucleotide Archive under accession number PRJEB105109.
Because of the strong correlations between atmospheric PM10 and atmospheric NOx and between tree circumference and tree height (Fig. 2), NOx and tree height were not included in the following analysis to avoid collinearity. A PERMANOVA indicated that all recorded properties (i.e. soil and environmental properties) contributed significantly to the bacterial community variation (Table 3). Principal coordinate analyses are visualized in Figs. S1–S4 in the Supplement. Soil pH was found to be the most influential factor for bacterial community composition among soil characteristics.
Table 3The explanatory power of different soil properties for the soil bacterial community variation according to a PERMANOVA (9999 permutations). SIRM served as a proxy for heavy metal pollution, trunk circumference served as a proxy for tree age, and air PM10 served as a proxy of traffic-related pollution.
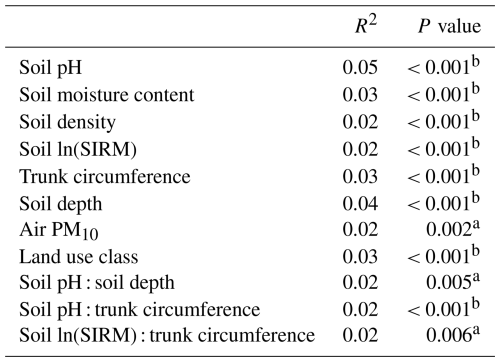
a p value <0.01,, b p value < 0.001.
Regarding richness indicators, we found that bacterial community diversity reached an optimum at a pH between 6 and 7 (Fig. 3a and b). Additionally, we found that soil moisture content was significantly correlated with ASV richness (p value = 0.047). Visualizing this trend indicated that diversity started to decrease more drastically as soil moisture content decreased (Fig. 3c and d). Site was also a significant factor for alpha diversity, but other soil characteristics and trunk circumference were not significantly correlated with soil ASV richness or the inverse Simpson index.
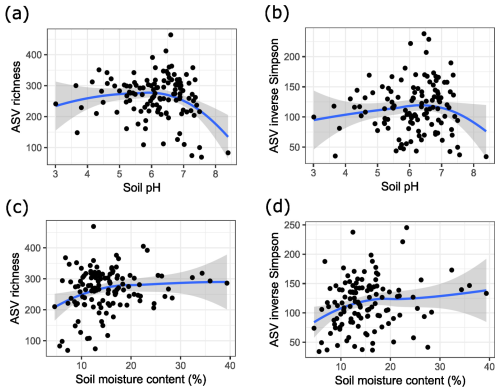
Figure 3(a) ASV richness of bacterial soil communities in relation to soil pH. (b) Diversity of bacterial soil communities expressed as the inverse Simpson index in relation to soil pH. (c) ASV richness of bacterial soil communities in relation to soil moisture content. (d) Diversity of bacterial soil communities expressed as the inverse Simpson index in relation to soil moisture content. The blue lines are the result of local polynomial regression fitting, and the grey area indicates the 95 % confidence interval.
To study the effect of tree circumference as a proxy for tree age, we used several approaches. We did not detect trends for co-occurring taxa clusters in trees with a similar tree circumference, which would have indicated ecological succession (Fig. S5). The soil communities at different sites, however, became less dissimilar as trunk circumference increased (p = 0.03, Kendall tau = −0.46), which indicates that bacterial communities converge as their associated vegetation becomes more established (Fig. S6). We also observed significant interaction effects between trunk circumference and other soil characteristics (Table 3). To further assess these interactions, we ran PERMANOVAs of four subsets of the data – small and large trunk circumference at two different soil depths. The results of these models confirmed that the potential impact of pH and SIRM on bacterial communities changed with trunk circumference: within soils of trees with larger trunk circumference (older trees) compared to those of trees with smaller trunk circumference (younger trees), we observed a clear decrease in the correlations with pH (R2 dropping from 0.19 in top soil and 0.23 at 10–15 cm depth to 0.09 and 0.11, respectively) and SIRM (R2 dropping from 0.14 in top soil and 0.14 at 10–15 cm depth to 0.05 and 0.04, respectively).
3.3 Bacterial communities at different spatial distance levels
A PERMANOVA to assess the effect of different spatial distance levels on the variation in the bacterial soil communities indicated that all levels contribute significantly to soil bacterial community variation (Table 4). Site, representing the largest spatial distances within this study, had the biggest impact on the community variation (R2 = 0.40). Two trees were selected per location, and we observed that tree ID additionally contributes to the community variation (R2 = 0.13). Variability of bacterial communities is further affected by the depth of the soil sample (within tree) and the sampling location at the surface as indicated by the orientation per tree (Fig. 2).
Table 4The PERMANOVA results (9999 permutations) for the effect of different spatial distance levels on the bacterial community. The factors are ordered from the largest to the smallest distance levels and align with the order in which they were added to the model.
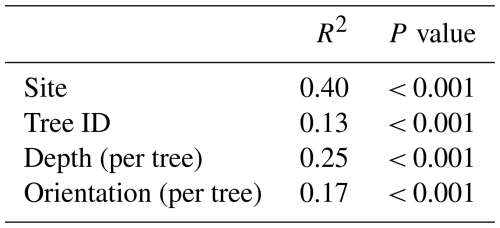
When adding the spatial distance level factors “site” and “tree ID” to the PERMANOVA of the soil characteristics (Table 3), we observed that some of the site variation is explained by soil characteristics (site goes from R2 = 0.40 to R2 = 0.30), but the effect of tree variation remains the same and is not further explained by soil characteristics.
In this exploratory study, 160 urban soil samples were collected between 13 February and 31 March 2023. As expected for urban soil, we found great heterogeneity of soil characteristics (Table 1) and bacterial communities (Table 4) in a single city. We could nevertheless identify some correlating factors for bacterial community variation, despite our single-time-point sampling approach. Within urban bacterial soil communities of this study, horizontal spatial distribution was found to be the most important driver of community variation. The importance of spatial distance became clear through the significantly large impact of site and tree ID as predictors of community variation, even when soil characteristics were considered covariables. This result suggests a strong effect by dispersal limitation on bacterial soil communities, in accordance with other soil studies (Bissett et al., 2010; Eisenlord et al., 2012; Richter-Heitmann et al., 2020). Additionally, as expected, we found that soil characteristics, land use, tree circumference (as a proxy for tree age), and pollution indicators also all significantly contributed to the bacterial community variation (Table 3).
Soil pH was identified as the most important explanatory variable of bacterial community composition among all soil characteristics (R2 = 0.05), which aligns with the results of Lauber et al. (2009) and Yan et al. (2016). In accordance with previous studies of natural soils by Fierer and Jackson (2006) and Lauber et al. (2009), we found the highest values of bacterial soil diversity at near-neutral pH (measured in KCl) (Fig. 3), suggesting an optimal pH for diverse communities. However, it is important to note that some low-diversity soils showed near-neutral pH too, highlighting the role of other factors shaping bacterial diversity alongside pH. Generally, bacterial communities in neutral soils have been observed to be more diverse and active than in acidic soils (Nugent and Allison, 2022). However, a more acidic soil pH may promote some desirable microbial functions such as increased carbon storage (Malik et al., 2018). While natural soils range in pH from acidic to neutral, urban soils are often alkalized (Nugent and Allison, 2022). In our study, the median pH was 6.4 (KCl), and we found that 93 % of the sampled soils had a higher pH than the median pH (KCl) of 4.6 of forest soils of a similar composition in the same region (De Keersmaeker et al., 2004). We also observed that soils in the “continuous urban fabric” land use type (> 80 % sealing) had a higher pH than those in the other land use types with less dense urban structures (Fig. 2c). Urban soil alkalinity is usually the consequence of leaching of calcareous substances from concrete, and an increased pH in soil has been associated with decreased microbial function (Nugent and Allison, 2022).
The gravimetric soil moisture content measured in this study was 15 % ± 7 %; however, these were single-time-point measurements which may have varied strongly with the weather conditions in the 47 d sampling period. Additionally, soil moisture content is known to correlate with organic matter content (Park et al., 1982), which we did not analyse in this study. This makes it difficult to compare to measurements of other studies, but we found that the literature reported on 21 % ± 2 % soil moisture content in a nearby region of agricultural, non-irrigated soil (Ryken et al., 2018). Previous studies reported typically drier soils in urban environments (Scalenghe and Marsan, 2009). Lower soil moisture content may have a great impact on urban soil ecology as anticipated by Nugent and Allison (2022). In this study, we observed a remarkable decrease in bacterial diversity in soils of lower moisture content (Fig. 3c and d). We hypothesize that soils in cities with a similar climate to the one addressed in this study (Antwerp, Belgium) may benefit from implementing measures to increase soil moisture content. Measures to increase soil moisture content include nature-based solutions, which decrease soil density and increase organic content, and technical solutions, such as removing soil sealing and draining, using permeable tiles, and improving water traps and storage (Scalenghe and Marsan, 2009; Zhang et al., 2021).
Trunk circumference of the tree under which the soil was sampled also had a minor but significant relationship with the bacterial soil community variation (R2 = 0.03). As the trees in our study were from the same species and grew as solitary trees in similar light conditions, it can be argued that tree circumference can be used as a proxy for the age of the tree and the extent of its root system (Van Mensel et al., 2023). However, tree growth rates in urban areas can vary because they depend on factors such as air pollution and soil compaction, which vary from site to site (Locosselli et al., 2019; Monteiro et al., 2017; Ruark et al., 1983). We could indeed not identify clear successional dynamics of the bacterial soil communities in this study using tree circumference as a time reference (Fig. S5) without actually sampling over time. Nevertheless, for bacterial community variation, we observed an interaction effect between the trunk circumference and soil pH and between the trunk circumference and the soil SIRM (Table 3). Soil SIRM is a proxy for heavy metal pollution in soil and is associated with concentrations of metals such as Fe, Pb, Cu, Zn, and Cd in urban roadside soils exposed to traffic pollution (Lu et al., 2011; Crosby et al., 2014). The effects of pH and heavy metal pollution on the community decrease as tree circumference increases, which suggests that over time, as root systems and soil bacterial communities develop, they stabilize and become less related to pH and heavy metal pollutants. Combined with the decrease in community dissimilarity over time, indicating a converging community composition (Fig. S6), this result implies that with tree development and/or time, bacterial communities in soils become more developed and, theoretically, more resilient. This is in line with the smaller potential impact of soil pH and metal pollution on the microbial community as the tree and likely its root system and associated soil microbiome develop. As it is not unusual for microbial soil communities to need several decades to develop after a disturbance (Zhou et al., 2017), more research is required to assess to what extent (repeated) disturbance of urban soils affects ecosystem service delivery and how we can better conserve older soils or speed up development of younger soils in the context of urban construction.
Other factors affecting urban soil communities included the soil density, soil sampling depth, modelled atmospheric pollution indicators (PM10), and land use class. The correlation between urban soil composition and air pollution could also reflect an association with nearby traffic intensity. Even artificial light at night has previously been shown to affect urban soil functioning (Li et al., 2023), but current research on these associations is very limited. Soil depth and density have previously been shown to be important drivers of soil microbial composition (Dick et al., 1988; Naylor et al., 2022). The bacterial community variation between different urban land use categories, however, has received less attention (Christel et al., 2023). Our study shows that the urban land use type, “roads and associated land”, “discontinuous urban fabric”, and “continuous urban fabric” (sealing > 80 %), around the sampled open soil also accounts for significant community variation, even after correcting for the correlations with pH, soil density, and pollution. This result indicates that future research is required to better understand the impacts of different urban land uses on urban soil functioning and its related ecosystem services.
Comparing R2 values of Tables 3 and 4, we find that spatial distance has a much greater impact on soil bacterial community variation than soil characteristics. Our results therefore indicate that the bacterial urban soil heterogeneity is not only caused by heterogeneity in soil characteristics (like pH and density), but also by other factors affecting site variability. This implies a strong effect by dispersal limitation on bacterial soil communities in accordance with other soil studies (Bissett et al., 2010; Eisenlord et al., 2012; Richter-Heitmann et al., 2020). We speculate that bacterial community variability is, in large part, caused by historic movement of soil and that the microbial inhabitants of the original source soil or exogenous replacing soil still affect the community composition of urban soils years later. Soil characteristics and potentially vegetation type and age put significant further selection pressure on the communities, but, in this study, the impact on community variation by soil pH is an order of magnitude smaller than the effect of site.
Finally, we believe it is noteworthy that, in this study, only the bacterial fraction of the soil microbiome was studied. For future research in urban soil ecosystem services, it is important to understand the composition and functions of fungi, archaea, protists, and eukaryotes, as well as those of bacteria, in urban soils.
To conclude, we observed that urban soil bacterial communities are indeed heterogeneous, which is likely a consequence of variation in the level and type of soil disturbance, e.g. by construction, other soil movement, and sealing practices. Nevertheless, just like in natural soils, soil characteristics such as moisture content and pH are correlated with soil bacterial diversity and composition. Our results support previous observations of more alkalized and drier urban soils. The fact that (i) bacterial urban soil communities are correlated with soil abiotic factors and (ii) bacterial community development is associated with the age of the vegetation and/or time since soil disturbance offers opportunities to contribute to soil diversity and ecosystem services through good soil management practices. This study therefore supports the importance of taking into account soil microbiome information when designing and implementing urban plans and implementing restoration strategies discussed by Schröder et al. (2024). However, without further knowledge on bacterial functioning and a proper definition of the characteristics of optimal bacterial communities in urban soils, our soil management suggestions remain hypothetical. Our study therefore highlights the need for future research to focus on understanding the required bacterial functions and compositions of “healthy” urban soils. As cities continue to expand, research on urban soil microbial communities is essential to develop sustainable urban planning and construction strategies that promote healthy soil ecosystems and overall urban wellbeing.
The underlying raw sequencing data are publicly available on the European Nucleotide Archive under accession number PRJEB105109 (https://www.ebi.ac.uk/ena/browser/view/PRJEB105109, last access: 27 January 2026).
The supplement related to this article is available online at https://doi.org/10.5194/we-26-35-2026-supplement.
KW and WS conceptualized the research, designed the methodology, and supervised the project. TB, NDG, ED, JD, and FG managed and executed the investigation. WS, TB, NDG, ED, JD, and FG curated the data. WS, TB, NDG, ED, JD, and FG executed the formal analyses. WS and ASL prepared the final paper based on the first draft prepared by TB, NDG, ED, JD, FG, and KW. All co-authors revised and edited the paper. KW did project administration, provided resources, and acquired the funding for this project.
The contact author has declared that none of the authors has any competing interests.
Publisher's note: Copernicus Publications remains neutral with regard to jurisdictional claims made in the text, published maps, institutional affiliations, or any other geographical representation in this paper. The authors bear the ultimate responsibility for providing appropriate place names. Views expressed in the text are those of the authors and do not necessarily reflect the views of the publisher.
The authors would like to thank the Centre for Medical Genetics Antwerp for the use of their Illumina MiSeq sequencing system. This publication has been prepared using the European Union's Copernicus Land Monitoring Service Information (Urban Atlas Land Cover/Land Use 2018).
This research has been supported by the Fonds Wetenschappelijk Onderzoek under the Research Foundation – Flanders (FWO; postdoctoral grant 12ZJ821N).
This paper was edited by Alexandra Rodríguez and reviewed by three anonymous referees.
Abrego, N., Crosier, B., Somervuo, P., Ivanova, N., Abrahamyan, A., Abdi, A., Hämäläinen, K., Junninen, K., Maunula, M., and Purhonen, J.: Fungal communities decline with urbanization – more in air than in soil, The ISME Journal, 14, 2806–2815, 2020.
Bainard, L. D., Klironomos, J. N., and Gordon, A. M.: The mycorrhizal status and colonization of 26 tree species growing in urban and rural environments, Mycorrhiza, 21, 91–96, 2011.
Bezberdaya, L., Kosheleva, N., Chernitsova, O., Lychagin, M., and Kasimov, N.: Pollution level, partition and spatial distribution of benzo (a) pyrene in urban soils, road dust and their PM10 fraction of health-resorts (Alushta, Yalta) and industrial (Sebastopol) cities of Crimea, Water, 14, 561 , 2022.
Bissett, A., Richardson, A. E., Baker, G., Wakelin, S., and Thrall, P. H.: Life history determines biogeographical patterns of soil bacterial communities over multiple spatial scales, Molecular Ecology, 19, 4315–4327, 2010.
Brockett, B. F., Prescott, C. E., and Grayston, S. J.: Soil moisture is the major factor influencing microbial community structure and enzyme activities across seven biogeoclimatic zones in western Canada, Soil Biology and Biochemistry, 44, 9–20, 2012.
Callahan, B. J., McMurdie, P. J., Rosen, M. J., Han, A. W., Johnson, A. J. A., and Holmes, S. P.: DADA2: high-resolution sample inference from Illumina amplicon data, Nature Methods, 13, 581–583, 2016.
Christel, A., Dequiedt, S., Chemidlin-Prevost-Bouré, N., Mercier, F., Tripied, J., Comment, G., Djemiel, C., Bargeot, L., Matagne, E., and Fougeron, A.: Urban land uses shape soil microbial abundance and diversity, Science of the Total Environment, 883, 163455, https://doi.org/10.1016/j.scitotenv.2023.163455, 2023.
City of Antwerp: Digital soil map city of Antwerp, https://portaal-stadantwerpen.opendata.arcgis.com/maps/386796b9907f4029a173bf33cfdd05c7 (last access: 27 January 2026), 2023.
Crosby, C., Fullen, M. A., and Booth, C. A.: Potential linkages between mineral magnetic measurements and urban roadside soil pollution (part 2), Environmental Science: Processes & Impacts, 16, 548–557, 2014.
De Keersmaeker, L., Martens, L., Verheyen, K., Hermy, M., De Schrijver, A., and Lust, N.: Impact of soil fertility and insolation on diversity of herbaceous woodland species colonizing afforestations in Muizen forest (Belgium), Forest Ecology and Management, 188, 291–304, 2004.
Delgado-Baquerizo, M., Eldridge, D. J., Liu, Y.-R., Sokoya, B., Wang, J.-T., Hu, H.-W., He, J.-Z., Bastida, F., Moreno, J. L., and Bamigboye, A. R.: Global homogenization of the structure and function in the soil microbiome of urban greenspaces, Science Advances, 7, eabg5809, https://doi.org/10.1126/sciadv.abg5809, 2021.
Dick, R., Myrold, D., and Kerle, E.: Microbial biomass and soil enzyme activities in compacted and rehabilitated skid trail soils, Soil Science Society of America Journal, 52, 512–516, 1988.
Eisenlord, S. D., Zak, D. R., and Upchurch, R. A.: Dispersal limitation and the assembly of soil Actinobacteria communities in a long-term chronosequence, Ecology and Evolution, 2, 538–549, 2012.
European Commission: EU Soil Strategy for 2030 – Reaping the benefits of healthy soils for people, food, nature and climate, https://eur-lex.europa.eu/legal-content/EN/TXT/PDF/?uri=CELEX:52021DC0699 (last access: 27 January 2026), 2021.
European Union's Copernicus Land Monitoring Service Information: Urban Atlas Land Cover/Land Use 2018 (vector), EEA [data set], https://doi.org/10.2909/fb4dffa1-6ceb-4cc0-8372-1ed354c285e6, 2020.
Fierer, N. and Jackson, R. B.: The diversity and biogeography of soil bacterial communities, Proceedings of the National Academy of Sciences, 103, 626–631, 2006.
Haahtela, T.: A biodiversity hypothesis, Allergy, 74, 1445–1456, 2019.
Hofman, J., Staelens, J., Cordell, R., Stroobants, C., Zikova, N., Hama, S., Wyche, K., Kos, G., Van Der Zee, S., and Smallbone, K.: Ultrafine particles in four European urban environments: Results from a new continuous long-term monitoring network, Atmospheric Environment, 136, 68–81, 2016.
Kuznetsova, A., Brockhoff, P. B., and Christensen, R. H. B.: lmerTest package: tests in linear mixed effects models, Journal of Statistical Software, 82, https://doi.org/10.18637/jss.v082.i13, 2017.
Lauber, C. L., Hamady, M., Knight, R., and Fierer, N.: Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale, Applied and Environmental Microbiology, 75, 5111–5120, 2009.
Lehmann, A. and Stahr, K.: Nature and significance of anthropogenic urban soils, Journal of Soils and Sediments, 7, 247–260, 2007.
Li, X.-M., Li, S., Huang, F.-Y., Wang, Z., Zhang, Z.-Y., Chen, S.-C., and Zhu, Y.-G.: Artificial light at night triggers negative impacts on nutrients cycling and plant health regulated by soil microbiome in urban ecosystems, Geoderma, 436, 116547, https://doi.org/10.1016/j.geoderma.2023.116547, 2023.
Locosselli, G. M., de Camargo, E. P., Moreira, T. C. L., Todesco, E., de Fátima Andrade, M., de André, C. D. S., de Andre, P. A., Singer, J. M., Ferreira, L. S., and Saldiva, P. H. N.: The role of air pollution and climate on the growth of urban trees, Science of the Total Environment, 666, 652–661, 2019.
Lovett, G. M., Traynor, M. M., Pouyat, R. V., Carreiro, M. M., Zhu, W.-X., and Baxter, J. W.: Atmospheric deposition to oak forests along an urban- rural gradient, Environmental Science & Technology, 34, 4294–4300, 2000.
Lu, S., Wang, H., and Guo, J.: Magnetic enhancement of urban roadside soils as a proxy of degree of pollution by traffic-related activities, Environmental Earth Sciences, 64, 359–371, 2011.
Malik, A. A., Puissant, J., Buckeridge, K. M., Goodall, T., Jehmlich, N., Chowdhury, S., Gweon, H. S., Peyton, J. M., Mason, K. E., and van Agtmaal, M.: Land use driven change in soil pH affects microbial carbon cycling processes, Nature Communications, 9, 3591, https://doi.org/10.1038/s41467-018-05980-1, 2018.
Mills, J. G., Weinstein, P., Gellie, N. J., Weyrich, L. S., Lowe, A. J., and Breed, M. F.: Urban habitat restoration provides a human health benefit through microbiome rewilding: the Microbiome Rewilding Hypothesis, Restoration Ecology, 25, 866–872, 2017.
Monteiro, M. V., Levanič, T., and Doick, K. J.: Growth rates of common urban trees in five cities in Great Britain: A dendrochronological evaluation with an emphasis on the impact of climate, Urban Forestry & Urban Greening, 22, 11–23, 2017.
Muyshondt, B., Wuyts, K., Van Mensel, A., Smets, W., Lebeer, S., Aleixo, C., Alós Ortí, M., Casanelles-Abella, J., Chiron, F., and Giacomo, P.: Phyllosphere bacterial communities in urban green areas throughout Europe relate to urban intensity, FEMS Microbiology Ecology, 98, fiac106, https://doi.org/10.1093/femsec/fiac106, 2022.
Naylor, D., McClure, R., and Jansson, J.: Trends in microbial community composition and function by soil depth, Microorganisms, 10, 540, https://doi.org/10.3390/microorganisms10030540, 2022.
Nugent, A. and Allison, S. D.: A framework for soil microbial ecology in urban ecosystems, Ecosphere, 13, e3968, https://doi.org/10.1002/ecs2.3968, 2022.
O'Riordan, R., Davies, J., Stevens, C., Quinton, J. N., and Boyko, C.: The ecosystem services of urban soils: A review, Geoderma, 395, 115076, https://doi.org/10.1016/j.geoderma.2021.115076, 2021.
Oksanen, J., Simpson, G., Blanchet, F., Kindt, R., Legendre, P., Minchin, P. R., O'Hara, R., Solymos, P., Stevens, M., Szoecs, E., Wagner, H., Barbour, M., Bedward, M., Bolker, B., Borcard, D., Carvalho, G., Chirico, M., De Caceres, M., Durand, S., Evangelista, H., FitzJohn, R., Friendly, M., Furneaux, B., Hannigan, G., Hill, M., Lahti, L., McGlinn, D., Ouelette, M., Ribeiro Cunha, E., Smith, T., Stier, A., Ter Braak, C., and Weedon, J.: vegan: community ecology package, R package version 2.6-4, https://CRAN.R-project.org/package=vegan (last access: 27 January 2026), 2022.
Park, H., Mok, S.-K., and Kim, K.-S.: Relationship between soil moisture, organic matter and plant growth in ginseng plantations, Journal of Korean Society of Soil Science and Fertilizer (Korea R.), 15, 1982.
R Core Development Team: R: A language and environment for statistical computing, R Foundation for Statistical Computing, Vienna, Austria, https://www.R-project.org/ (last access: 27 January 2026), 2006.
Richter-Heitmann, T., Hofner, B., Krah, F.-S., Sikorski, J., Wüst, P. K., Bunk, B., Huang, S., Regan, K. M., Berner, D., and Boeddinghaus, R. S.: Stochastic dispersal rather than deterministic selection explains the spatio-temporal distribution of soil bacteria in a temperate grassland, Frontiers in Microbiology, 11, 1391, https://doi.org/10.3389/fmicb.2020.01391, 2020.
Ruark, G., Mader, D., Veneman, P., and Tattar, T.: Soil factors related to urban sugar maple decline, Arboriculture & Urban Forestry (AUF), 9, 1–6, 1983.
Ryken, N., Nest, T. V., Al-Barri, B., Blake, W., Taylor, A., Bodé, S., Ruysschaert, G., Boeckx, P., and Verdoodt, A.: Soil erosion rates under different tillage practices in central Belgium: New perspectives from a combined approach of rainfall simulations and 7Be measurements, Soil and Tillage Research, 179, 29–37, 2018.
Scalenghe, R. and Marsan, F. A.: The anthropogenic sealing of soils in urban areas, Landscape and Urban Planning, 90, 1–10, 2009.
Schröder, A., Schloter, M., Roccotiello, E., Weisser, W. W., and Schulz, S.: Improving ecosystem services of urban soils–how to manage the microbiome of Technosols?, Frontiers in Environmental Science, 12, 1460099, https://doi.org/10.3389/fenvs.2024.1460099, 2024.
Shade, A., McManus, P. S., and Handelsman, J.: Unexpected diversity during community succession in the apple flower microbiome, mBio, 4, e00602–00612, https://doi.org/10.1128/mBio.00602-12, 2013.
Smets, W., Spada, L. M., Gandolfi, I., Wuyts, K., Legein, M., Muyshondt, B., Samson, R., Franzetti, A., and Lebeer, S.: Bacterial succession and community dynamics of the emerging leaf phyllosphere in spring, Microbiology Spectrum, 10, e02420–02421, https://doi.org/10.1128/spectrum.02420-21, 2022.
Van Mensel, A., Wuyts, K., Pinho, P., Muyshondt, B., Aleixo, C., Orti, M. A., Casanelles-Abella, J., Chiron, F., Hallikma, T., and Laanisto, L.: The magnetic signal from trunk bark of urban trees catches the variation in particulate matter exposure within and across six European cities, Environmental Science and Pollution Research, 30, 50883–50895, 2023.
Vu, Q., Marwick, B., Hester, J., and Held, M.: Package `ggbiplot', https://doi.org/10.32614/CRAN.package.ggbiplot, 2011.
Wang, Y.-P. and Goll, D. S.: Modelling of land nutrient cycles: recent progress and future development, Faculty Reviews, 10, https://doi.org/10.12703/r/10-53, 2021.
Wickham, H.: ggplot2: elegant graphics for data analysis, Springer, ISBN 978-3-319-24275-0, 2016.
Wittouck, S., Van Rillaer, T., Smets, W., and Lebeer, S.: Tidytacos: An R package for analyses on taxonomic composition of microbial communities, Journal of Open Source Software, 10, 6313, https://doi.org/10.21105/joss.06313, 2025.
Wuyts, K., De Schrijver, A., Staelens, J., and Verheyen, K.: Edge effects on soil acidification in forests on sandy soils under high deposition load, Water, Air, & Soil Pollution, 224, 1–14, 2013.
Wuyts, K., Smets, W., Lebeer, S., and Samson, R.: Green infrastructure and atmospheric pollution shape diversity and composition of phyllosphere bacterial communities in an urban landscape, FEMS Microbiology Ecology, 96, fiz173, https://doi.org/10.1093/femsec/fiz173, 2020.
Yan, B., Li, J., Xiao, N., Qi, Y., Fu, G., Liu, G., and Qiao, M.: Urban-development-induced changes in the diversity and composition of the soil bacterial community in Beijing, Scientific Reports, 6, 38811, https://doi.org/10.1038/srep38811, 2016.
Yu, H., Li, T., Liu, Y., and Ma, L.: Spatial distribution of polycyclic aromatic hydrocarbon contamination in urban soil of China, Chemosphere, 230, 498–509, 2019.
Yuan, X., Xue, N., and Han, Z.: A meta-analysis of heavy metals pollution in farmland and urban soils in China over the past 20 years, Journal of Environmental Sciences, 101, 217–226, 2021.
Zhang, Y., Qiao, L., Chen, C., Tian, L., and Zheng, X.: Effects of organic ground covers on soil moisture content of urban green spaces in semi-humid areas of China, Alexandria Engineering Journal, 60, 251–259, 2021.
Zhou, Z., Wang, C., Jiang, L., and Luo, Y.: Trends in soil microbial communities during secondary succession, Soil Biology and Biochemistry, 115, 92–99, 2017.





